Correlated Genes¶
Overview¶
The Correlated Genes Tab shows the genes that are either up or down-regulated with respect to selected gene in the Feature View. The values are based on the public transcriptomics data that are available in BV-BRC.
See also¶
Examining Transcriptomics Data Tutorial
Most of the Ptranscriptomics data have been curated from published gene expression datasets related to bacterial pathogens in NCBI’s GEO database. Some additional data sets have been incorporated from the NIAID-funded [Systems Biology and Functional Genomics Centers and other sources.
Accessing the Correleated Genes¶
Clicking the Correlated Genes tab in the Feature View displays a table showing all the genes with expression data that are either up or down-regulated with respect to selected gene.
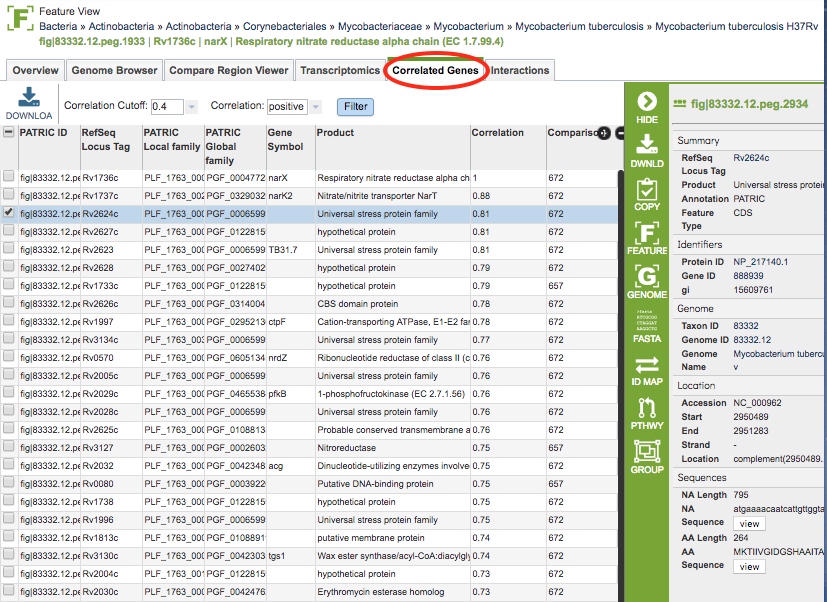
Correlated Genes Table Tools¶
You may do the following with the table:
Download the entire contents of the data used to create the charts in text, CSV, or Excel format by clicking the Download button above the table on the left side.
Filter the correlated genes based on positive or negative correlation and cutoff value using the dropdown boxes above the table and clicking the Filter button.
The columns in the table include the following:
ID: Unique identifier for the gene, usually of the form fig|#####.##.peg.####
RefSeq Locus Tag: RefSeq locus tag for the gene, if available
Local Family: “Local” Protein Family containing the gene
Global Family: “Global” Protein Family containing the gene
Gene Symbol: Gene symbol, if available
Product: Gene product, as annotated by BV-BRC
Correlation: Correlation value of the gene compared to the gene in the Feature View
Comparisons:
Action buttons¶
After selecting one or more of the experiments by clicking the checkbox beside the Title column in the table, a set of options becomes available in the vertical green Action Bar on the right side of the table. These include
Hide/Show: Toggles (hides) the right-hand side Details Pane.
Download: Downloads the selected items (rows).
Copy: Copies the selected items to the clipboard.
Feature: Loads the Feature Page for the selected feature. Available only if a single feature is selected.
Features: Loads the Features Table for the selected features. Available only if multiple features are selected.
Genome: Loads the Genome View Overview page corresponding to the selected feature. Available only if a single feature is selected.
Genomes: Loads the Genomes Table, listing the genomes that correspond to the selected features. Available only if multiple features are selected.
FASTA: Provides the FASTA DNA or protein sequence for the selected feature(s).
ID Map: Provides the option to map the selected feature(s) to multiple other idenfiers, such as RefSeq and UniProt.
MSA: Launches the Multiple Sequence Alignment (MSA) tool and aligns the selected features by DNA or protein sequence in an interactive viewer.
Pathway: Loads the Pathway Summary Table containing a list of all the pathways in which the selected features are found.
Group: Opens a pop-up window to enable adding the selected sequences to an existing or new group in the private workspace.