Speciality Genes¶
Overview¶
The Specialty Genes Tab provides a table of all the annotated “specialty genes” (virulence factors, antibiotic resistance genes, drug targets, human homologs, transporters, and essential genes) corresponding to the set of genomes in the selected Taxon View level or in the user-defined Genome Group. From this page, features can be sorted, filtered, collected into groups, and downloaded.
See also¶
Accessing the Specialty Genes Table¶
Clicking the Specialty Genes Tab in a Taxon View displays the Specialty table (shown below), listing all the features annotated as Specialty Genes corresponding to the set of genomes in the selected taxon level.
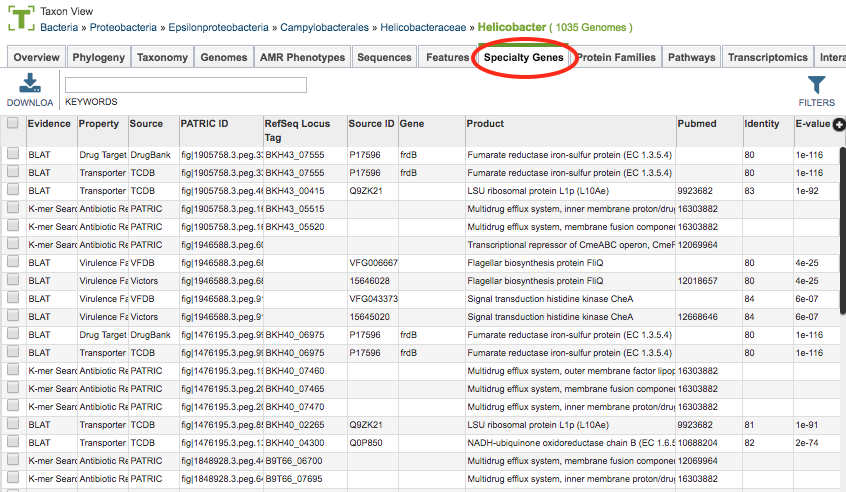 BV-BRC identifies features as Specialty Genes via BLASTP-based sequence similarity mapping to reference genes are collected from reputed external databases or manually curated by the BV-BRC team. These features (genes) include
BV-BRC identifies features as Specialty Genes via BLASTP-based sequence similarity mapping to reference genes are collected from reputed external databases or manually curated by the BV-BRC team. These features (genes) include
Antibiotic Resistance: Mapped from CARD, NDARO, and curated AMR genes.
Human Homologs: Mapped from Reference Human Genome at NCBI.
Virulence Factors: Mapped from VFDB, Victors, and curated virulence factors.
Transporters: Mapped from TCDB.
Essential Genes: Mapped to essential genes (using flux-balance for Reference and Representative genomes, predicted using flux-balance analysis.
See Specialty Genes for additional information.
Specialty Gene Table Tools¶
Within this table you may do the following:
Download the entire contents of the table in text, CSV, or Excel format by clicking the Download button above the table on the left side.
Rearrange and narrow the list of sequences in the table via sorting (using column headers), keywords (using the Keyword box), and filtering (using the Filters tool).
Filter Tool¶
As with all tables, the Filters tool is available to narrow the display of the items in the table, show below:
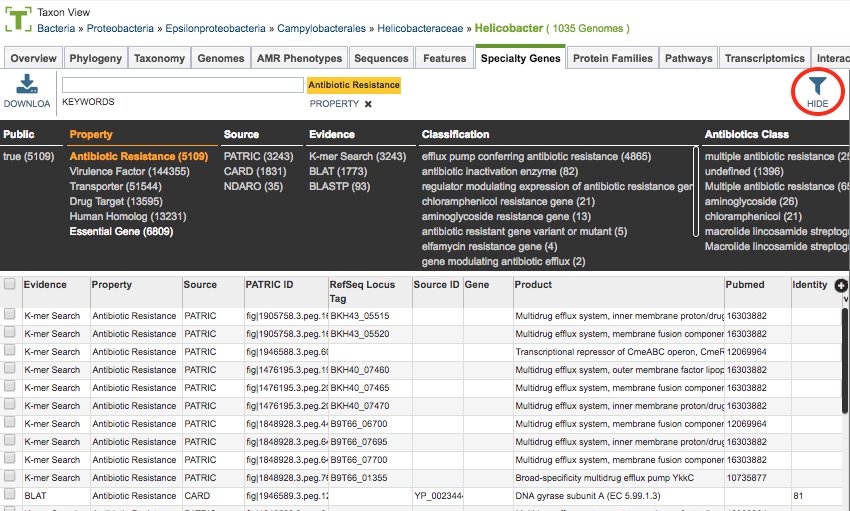
Clicking on the Filters button at the top right of the table opens the Filter Panel above the table, displaying column names from the table and values for those columns with counts of occurence. Clicking on the filter values narrows the features displayed in the table to those matching the chosen filter values. Clicking the Hide button closes the Filter Panel. More details are available in the Tables and Filters .